|
Back to Blog
Antibody cross reactivity7/8/2023 Understanding cross-reactive immune responses to SARS-CoV-2 may contribute to our understanding of the heterogeneity of clinical outcomes in COVID-19 disease and vaccination and allows us to identify immune epitopes that are conserved between genetically diverse coronaviruses and are most likely to be functionally constrained. But in other cases, such as dengue virus and zika virus infections 36, 37, 38, disease severity is potentially exacerbated (Box 1). 1), in some cases promoting protection against infections - such as in influenza A 34 and Japanese encephalitis virus infections - in which beneficial cross-reactive T cell responses are found 35. Evidence from the study of other infectious diseases suggests that pre-existing, cross-reactive immune response may impact disease outcomes (Fig. However, the exact source and the subsequent role of pre-existing SARS-CoV-2 cross-reactivity remain a question of considerable research interest. Given the high homology of SARS-CoV-2 with other HCoVs, particularly those of the β-CoV genus, and the very high prevalence of CCCs, it is probable that these viruses are the source of cross-reactive immune responses. Adaptive immune responses, including the development of a coordinated SARS-CoV-2-specific CD4 + and CD8 + T cell response and neutralizing antibodies, are associated with a reduction in COVID-19 disease severity 20, 21, 22 however, certain T cell responses including unconventional CD16 + T cells 23, mucosal-associated invariant T (MAIT) cells 24 and γδ T cells 25 may be increased in severe COVID-19 and are potentially related to immunopathology.Ĭross-reactive immune responses that exist in individuals before SARS-CoV-2 exposure may also affect susceptibility to infection and disease severity 4, 26, 27, 28, 29, 30, 31, 32, 33. Elevated serum cytokine levels are associated with severe COVID-19 infections and are a strong predictor of adverse disease outcomes 19. These include autoantibodies to type I interferons 15, 16 altered myeloid cell populations including increased numbers of immature neutrophils and loss of non-classical monocytes 17 as well as increased hyperinflammatory and aberrant CD163 + monocytes 18. Moreover, various immunological correlates have been associated with COVID-19 severity and disease outcomes. These include demographic factors such as age 9, Black and Asian ethnicity 10, male gender 11 and comorbidities such as diabetes 12, kidney disease, cerebrovascular disease, cardiovascular disease and respiratory disease 13, 14. Several risk factors for COVID-19 disease severity have been identified. Although SARS-CoV-2 is broadly classified as pathogenic owing to its relatively high case–fatality ratio 7, COVID-19 shows a diverse range of symptoms from asymptomatic or mild disease in the majority of patients, to critical disease including acute respiratory distress syndrome, pneumonia, cardiac arrythmia, encephalopathy, acute kidney injury and, in some cases, death 8. By contrast, MERS-CoV, SARS-CoV and SARS-CoV-2 have all emerged in the past 20 years and can cause severe, potentially fatal disease 6. The so-called common cold coronaviruses (CCCs) HKU1, OC43, NL63 and 299E are seasonal, are common in human populations globally and were responsible for an estimated 10–20% of viral respiratory infections in 2019, causing predominantly mild disease 5. There is a high degree of homology among HCoVs in both structural proteins and NSPs 3, 4. 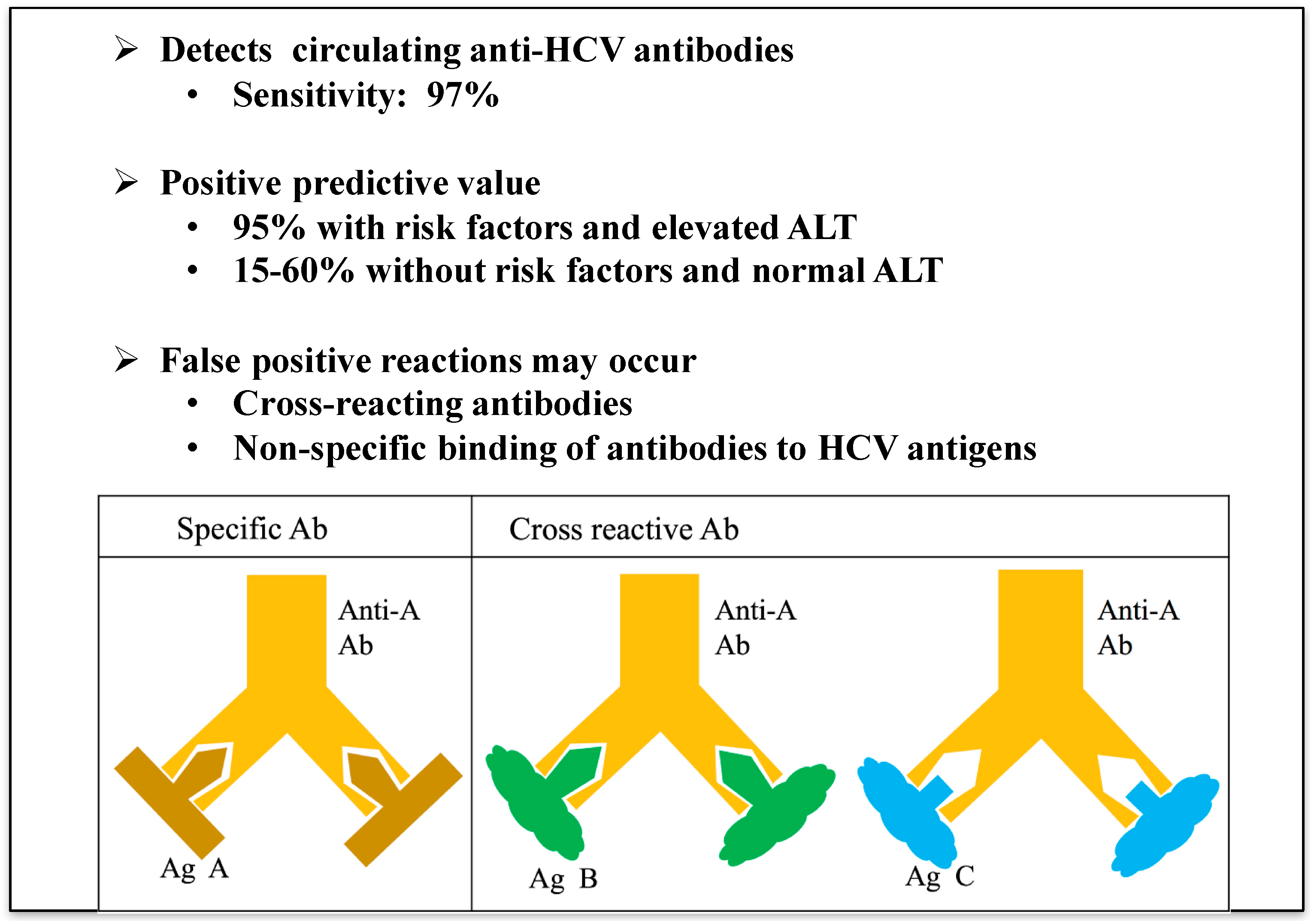
All HCoVs express the structural proteins spike, envelope, membrane and nucleocapsid, accessory proteins in open reading frames (ORFs) 3, 6, 7, 8, 9, 10 and 14, the much larger group of non-structural proteins (NSPs) in ORF1 (ref. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causal agent for COVID-19, is a member of a large family of positive-sense RNA human coronaviruses (HCoVs), which include the Alphacoronaviruses (α-CoVs) 299E and NL63 and the Betacoronaviruses (β-CoVs) HKU1, OC43, Middle East respiratory syndrome coronavirus (MERS-CoV), SARS-CoV and SARS-CoV-2 (ref.
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
